GIMS.dx
What is GIMS.dx?
GIMS.dx is a platform to improve processes of any NGS-based molecular diagnostic: germline and somatic. This includes genetic diagnostics of rare hereditary diseases, cancer predispositions and sequencing analyses of tumor materials as well.
The core feature of GIMS.dx is our recently developed DNAwiz, that harbours structured step by step decision-making and doumentation-workflows for variant classification, clinical interpretation and recommendation.
With GIMS.dx physicians and scientists in genetics and pathology can do their important work in a structured ecosystem that is the basis for better medical decisions.
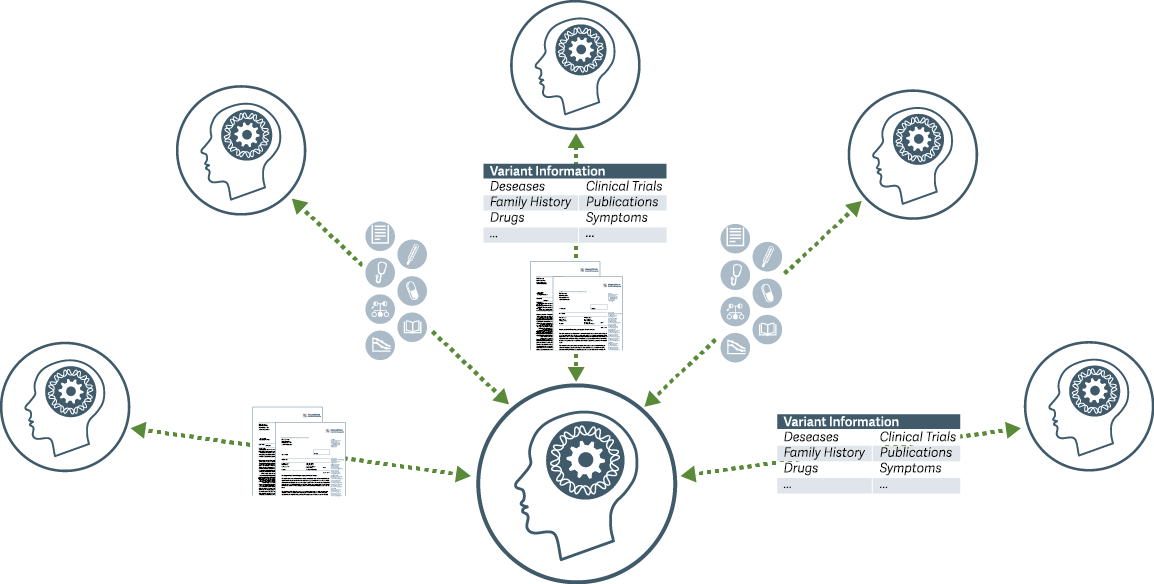
By using GIMS, molecular and clinical teams work together in a digital genetic ecosystem. The aggregated knowledge and findings become transparently trusted and disseminated: the fair sharing of variant information, genetic knowledge and clinical observations is part of a continuous truth-finding process.
How DNAwiz works

- variant list (vcf) is uploaded with standard interfaces or manually
- if a variant is already available in the system and trustfully classified, it can be processed automatically for generation of a report in clinical grade
- to process formerly not classified variants the user is guided through the variant assessment and interpretation process step by step. Decision and argumentation processes can be document and versionised
- where possible, the user can also draw variant information and existing assessments from the sharing network
- Completed assessments can be reviewed and approved using customizable workflows
- Interpretations, diagnosis and recommendations are compiled in a clinical report available in multiple formats using content from the database amended with the user´s comments where applicable
- Results can be
- provided to physicians and patients as printed documents
- incorporated into EHR systems or
- distributed using customizable digital channels
- once assessments has reached a level of maturity, the customer can choose to share the knowledge within the network
- detected variants are listed and automatically connected with metadata from public sources
Key benefits
Stepwise guidance through variant classification and interpretation process:
- identified variants automatically equipped with available metadata from public databases (e.g. ref genome, gene no., transcript. no., chromosome, rs no., variant information)
- increased productivity, decrease in time required for variant analysis
Structured documentation process with clear-cut workflows for review and approval processes:
- makes decision processes transparent and reproducible – ideal also for QA and audits
- provides curated information for diagnostic decision support
- allows structured compilation of own knowledge
- supports automated production of diagnostic reports
- enables time savings and ensures standardization and constant quality of diagnostic output
- enables sharing of content and knowledge within a growing network of laboratories, scientific institutions and expert groups
Contact block
PLEASE NOTE: By filling out the contact form and activating the send button you consent to the processing of the stated data according to art. 6, para. 1a EU-GDPR for the purpose of processing your inquiry. More information can be found in our privacy policy.
